Throwing Light on Throwing Light on Machine Learning
Here is a “making of the paper” post on my recent Data Science Journal article on transparency in Machine Learning, a case study in how journals can handle the meteoric rise in the use of AI-methods.
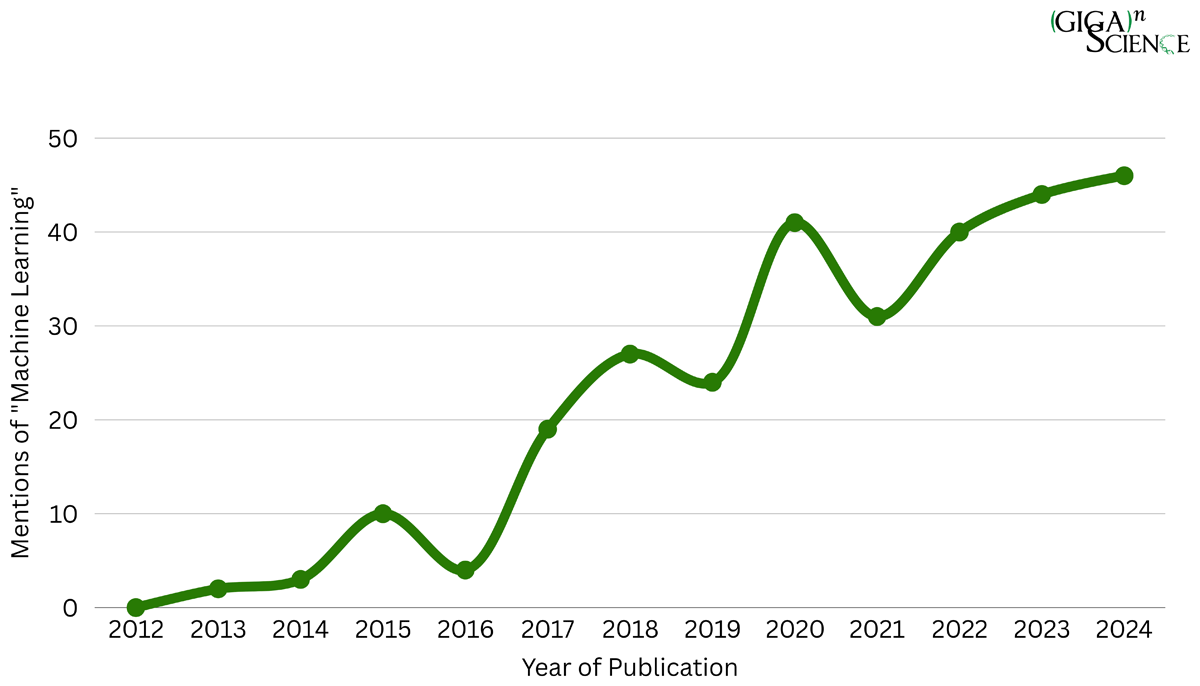
With the meteoric rise in the use of AI-methods in scientific research, how can journals and peer reviewers continue to review and assess science in a reproducible, transparent and trustworthy manner? AI-based approaches such as Machine Learning (ML) are extremely powerful tools for carrying out research, but through their complexity and “black box” and often proprietary (non-open source) nature, the intelligibility of these methods can be even more challenging to comprehend than traditional computational methods. Last month my paper in Data Science Journal presented a case study trying to address this very challenge, and as I promised when it came out to write up a more detailed “making of the paper” post I am finally providing some more behind-the-scenes insight here. This paper was written from the perspective of a journal focusing on publishing data intensive computational research in a reproducible and transparent manner, that was now struggling to maintain this track record with an increasing proportion of submissions using ML. As you can see in Figure 2 of the paper, looking at GigaScience publications using the phrase ‘machine learning’ since the launch of the journal in 2012 you can see a steady rise in ML content. This trend is mirrored in the scientific literature as a whole, in 2024 alone PubMed published a total of 43,931 articles covering ‘machine learning’.
Running GigaScience this was the situation we found ourselves at the start of the decade, having to constantly tailor (make up as we went along) the review processes and editorial thresholds for this type of work. We’d experimented using platforms such as Gigantum to wrap and showcase the reproducibility of ML research, but using third-party cloud-based tools wasn’t necessary a scalable and stable approach to tackle the rapid rise of this research (as demonstrated by the Gigantum platform closing down a few years later). In June 2021 Jen Harrow invited me to a workshop of the ELIXIR machine learning Focus Group bringing together different stakeholders together to discuss the DOME Recommendations for Machine Learning research they had just developed. DOME standing for “Data, Optimization, Model and Evaluation” as each component of a ML implementation usually falls within one of these four topics. Capitalising on the acceptance of their DOME-ML guidelines in a special issue of Nature Methods on “Reporting standards for machine learning” that was coming out later that year, the aim of the workshop was to discuss how the DOME Recommendations could be adopted by relevant journals, publishers and funders, and also to start a discussion for a process towards a governance structure around these recommendations. Being based in Asia it wasn’t practical for me to attend the workshop at the time, but our Data Scientist Chris Armit was keen to get involved so we made him our point person with the ELXIR ML community, and he participated and represented us in their ongoing working group meetings until his departure in summer 2024. The DOME recommendations and the other guidelines published in the Nature Methods special issue were immediately useful in how we handled the peer review of ML-research, as we finally had a logical framework and standards to work from as we assessed this research.
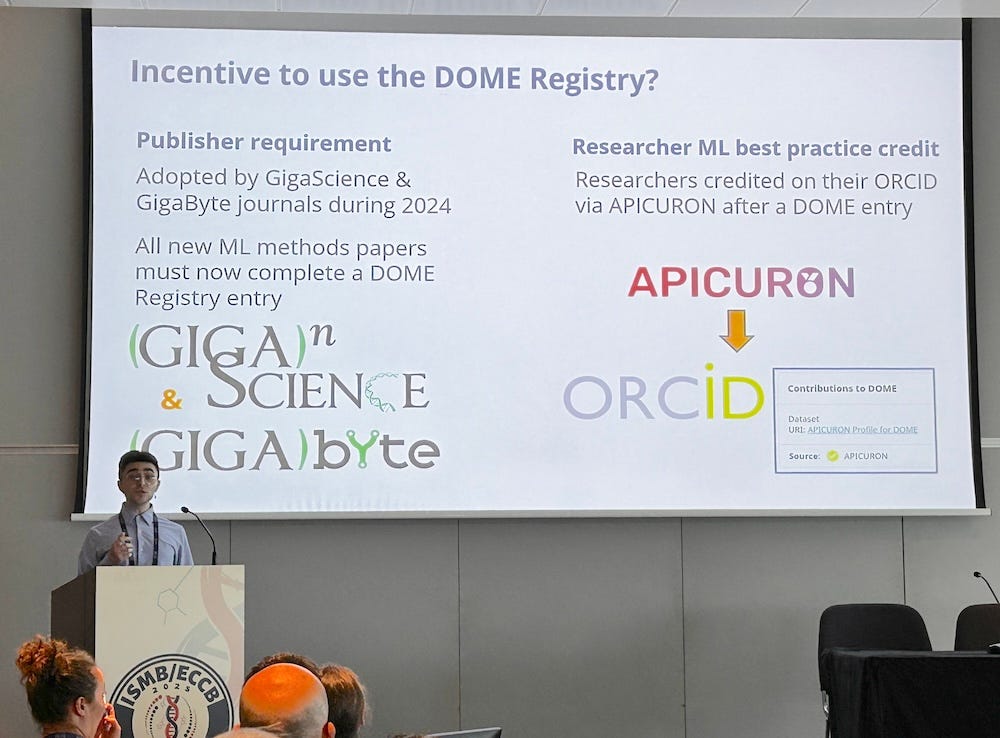
Chris Armit presenting our DOME trial at the BOSC 2024 meeting in Montreal.
DOMEeability and the “No DOME, Go Home” Standard
Alongside DOME-ML, the special issue also published the AIMe registry for AI in bioMedical research, alongside the “Bronze, Silver, Gold” reproducibility standards for machine learning in the life sciences from Benjamin Heil et al. We found the Heil et al. standards particularly useful for editorial triage, as we could use the Bronze standard (a minimal standard for reproducibility where the data, models and code used in the analysis need to be publicly available) as an easy minimum criteria to send papers out to peer review, and then aiming at least for their Silver standard as a criteria for publication (having additional minimal methodological details provided and the dependencies of the analysis able to be downloaded and installed in a single command). On top of working directly with the DOME community we also discussed these issues with some of the Heil et al. authors, particularly Casey Greene who joined our Editorial Board and was a very useful sounding board in developing our ML policies and workflows.
Using a combination of the “Bronze, Silver, Gold” approach alongside the required fields in the DOME reproducibility standards we quickly developed an in-house protocol on whether to send ML submissions to review. If these manuscripts were close enough to compliance that any missing minor methodological details could likely be added during curation and review we would give them a chance. But if there was likely no chance of ever meeting the DOME standards we could save everyone involved a lot of time and effort and immediately reject these submissions. In our triage meetings the phrase “no DOME, go home” became a bit of an inside joke amongst the editors for describing the most unreproducible papers (also helping exclude any ghost written and AI-generated papers from paper mills that immediately fail these tests through not having any real data, models or evaluations to back up the writing). The GigaScience curation team also finding these guidelines very useful to work into their curation workflows and practices as well.
Having the journal directly involved in the ELIXIR ML Focus Group meetings was very useful as they wanted to work towards a list of recommendations that are more tailored to the needs of a journal/publisher, and we could also provide feedback and help customize the curation tools and registry they were developing so we could integrate these more smoothly into our review processes. The PI’s of the project Silvio Tosatto and Fotis Psomopoulos were very welcoming and helpful, and also we had a lot of useful interactions with Alex Miguel Monzon at our birthday parties at ISMB. We also have to give very special thanks the very responsive development and curation team, particularly Gavin Farrell and Omar Attafi.
The Wonderful Wizard of ML
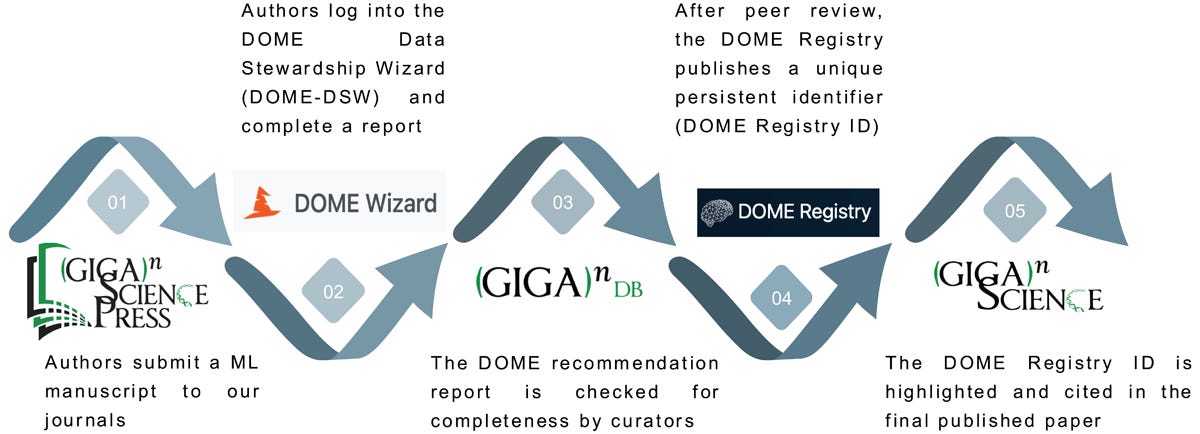
On top of using the DOME recommendations in Editorial triage our curation team started collecting annotation information on these ML experiments, but to start with there was no easy mechanism by which authors could submit their DOME recommendations to the journal. In April 2023 Chris Armit presented our early efforts gathering this information at the DOME Strategic Implementation Study KO meeting in Padova, and one of the outcomes of this was DOME then provided us with the option for researchers to share DOME annotations using the DOME Data Stewardship Wizard tool. We initially started collecting these annotations in a wordfile published alongside the dataset in GigaDB, but once the wizard came on line we included the links to the DOME wizard in the GigaDB dataset. You can see the first example of a published DOME annotation in March 2023, where the GigaDB dataset had the annotations included as a word file (and the link to the registry was then added in later). Working with the DOME developers we then received more stable DOME wizard links, so we worked these DOME annotations into the peer review process. For the DOMEeable ML papers we decided to send out to review, before peer review our curation team would send the DOME wizard link to the authors to provide details into all of the required fields. There could be some back and forth to obtain all of this information, making sure the input data and models were publicly available, and if necessary updating the paper to make sure these details were included for scrutiny. When sufficient information was available then we would then send the paper to peer review, and we make sure the peer reviewers received the link to the DOME annotations so they could scrutinize these. Rather than providing additional work for the reviewers, we hoped that presenting this key methodological information in a more transparent and structured manner should actually make the peer review process potentially easier and quicker as the reviewers would save time having to dig all this crucial information out.
Summary of the steps taken that utilise the DOME guidelines, DOME-DSW, and DOME Registry to aid the peer review of ML research in our final workflow.
Throughout 2023 and 2024 we continued to test and fine tune this processes as the number of ML submissions continued to rise. Going beyond including the annotations alongside the dataset, we started highlighting the annotation links in the published papers so they would be more prominent and discoverable for readers. As with reviewers, we thought providing this key methodological information in a more transparent and structured manner would be useful for downstream users who may only be interested in finding useful training data, reusable models, or scrutinizing specific methodological details. At the initial workshop Chris attended in June 2021 there was talk of a DOME registry being developed, and in September 2022 a rudimentary version of this was launched. Our annotations started being transferred into this as 2023 and 2024 progressed but this was a slow manual process and wasn’t coordinated with publication. Working with the DOME Registry developers our curation team eventually managed to streamline this process, getting permission to update and curate the registry details themselves, and eventually allowing us to be able to create the DOME registry entries. Allowing us to follow the best practices of the Data Citation Principles and cite the DOME registry PIDs in the references of the papers. In December 2024 the DOME Registry paper was published in GigaScience, and it was nice we managed to showcase our now fine-tuned and optimised publisher case study in the paper.
Read All About It
As publisher engagement was an important project deliverable for the ELIXIR machine learning Focus Group they were very keen to share our experiences and promote the potential of DOME tools and standards for other journals to address the problem of poor quality unreproducible ML research being published. On top of presenting our case study in a couple of ELIXIR webinars, they also produced leaflets and other materials promoting our example.
In July 2024 Chris Armit left GigaScience Press but just before his departure he presented a preliminary version of our case study at the BOSC 2024 meeting in Montreal (see the video of his talk).
As part of his handover he wrote up a first draft of this paper, written very much from the perspective of how the GigaScience team is currently handling this specific workflow. Yannan Fan from our curation team took over the handling of these ML papers and started attending the regular ELIXIR machine learning Focus Group calls, and Chris Hunter and I also joined some of the calls and helped with this continued collaboration. Chris Hunter joined the SAB, and in October 2025 the DOME activities moved to the ELIXIR AI Ecosystems Focus Group.
Assisting with this outreach I also promoted DOME at BGI’s ICG conference and the OUP Computational Biology Senior Editors’ Forum. Submitting an abstract to the International Data Week (IDW 2025) Conference in Brisbane in October this seemed to be a topic of interest as it was accepted for a full talk, so I thought I would use that opportunity to complete the paper and submit it to the accompanying International Data Week collection at Data Science Journal. As we now had published over 50 examples using this workflow it felt like a good opportunity to look at the lessons learned. With the tools and curation processes ironed out the workflow finally felt quite mature it, so it felt useful to share this approach much wider so others could potentially adapt and follow it. After the GigaScience Press and DOME teams had put so much work in developing this workflow, sharing the technical details more widely to allow replication also felt a responsible thing to do so the institutional knowledge developed to run this workflow wasn’t just confined to just one journal. Which in retrospect was wise considering shortly after submission of the paper most of the team working on this project were suddenly fired by BGI.
Updating and completing the paper, I broadened it to include a mapping of the current landscape of ML standards as there were a lot more examples than we were initially aware of. There is a huge number of field-specific ML standards, particularly in medical AI, where at least 26 reporting guidelines were published between 2009 and 2023. As a multi-disciplinary journal with a very broad scope we needed to use more general, domain-agnostic standards, but on top of the three standards published in the Nature Methods special issue, in 2024 the REFORMS consensus-based recommendations for ML-based science was also published in this more generalist space. Our initial reason for testing DOME was because the DOME community approached us, but working on this paper provided a useful opportunity to assess newer standards that were not developed when we started working with DOME, and also go back and better assess the alternative standards that we may have missed at the time. Our paper presents this mapping and comparisons between the more general ML standards, and it was interesting to see where DOME sat in this space, being broad but not as overly complicated as some of the other standards. On top of the complexity issue, the extremely active and helpful community, alongside the ease of use of the wizard and registry were probably the main selling points for us working with DOME. The other generalist standard communities lacking in this area, which can be seen in the continued growth and development of the DOME registry .
Sending the paper to peer-review in Data Science Journal, the reviewers were generally positive but requested more systematic comparative analysis and researcher feedback. Collecting more examples and a longer time period to gather more quantitative data such as citations would have been necessary to provide statistical significance for such comparative analyses, but our rationale for writing this paper was to present a n=55 case study from a 3 year pilot that others could learn from. Using this effort as the start of a longitudinal study it would be very interesting to come back when more data has been collected and try to do these types of comparative analyses in a follow-up paper. We had very limited time to address the revision deadlines, but Gavin Farrell from DOME was extremely helpful in providing us with some of their usage analytics and insight into their curation times to help address the comments.
Gavin Farrell presenting the DOME Registry (and our collaboration) at BOSC2025 in Liverpool.
We also sent out a quick anonymous survey to all of the corresponding authors of these paper to try to get some feedback and insight into how much effort and time it took to fill out the DOME wizard, and generally the responses were positive. We included this additional data, and speculated a bit about the time costs and this was then sufficient to get the paper accepted for publication. I have a lot more to say on the cost v benefit issues this raised but I think I will save that for a follow up post as I’ve already written too much here.
Please check out the final published paper or get in touch if you have any questions on any of this. On top of the main IDW talk I was also asked to present a shorter version of the cases study in the “Bridging the FAIR gap: transforming the long tail of supplementary data & generalist repositories into FAIR datasets” session of IDW, and the video of this tl;dr version is available here.
Thank You For The ML-sic
For this to have come together I need to thank the GigaScience curation (particularly Chris Armit, Chris Hunter and Yannan Fan who worked most closely with the DOME team) and Editorial teams (Nicole Nogoy, Hongling Zhou, Hongfang Zhang and Hans Zauner). The DOME community has been particularly welcoming and helpful, and I need to acknowledge Jen Harrow, Silvio Tosatto, Fotis Psomopoulos, and Alex Miguel Monzon for getting us involved in the efforts of the ELIXIR machine learning Focus Group, and Gavin Farrell and Omar Attafi who really helped us on the curation side and in getting the paper out. Gavin providing us with lots of useful information for the paper, and Omar and Yannan tidying up and updating all of the metadata for our 55 submissions in the DOME registry by the time of publication. Our summer intern Chima Okafor from HKUST needs to be acknowledged for a great job helping with the some of the figures. With final thanks also to CODATA and their Data Science Journal for reviewing and accepting the conference and journal submissions. You can see more on IDW2025 in my write-up here.
Further Reading
Edmunds, S.C., Nogoy, N., Lan, Q., Zhang, H., Fan, Y., Zhou, H. and Armit, C. Integrating Machine Learning Standards in Disseminating Machine Learning Research. Data Science Journal, 25(1) (2026). https://doi.org/10.5334/dsj-2026-001
Walsh, I., Fishman, D., Garcia-Gasulla, D. et al. DOME: recommendations for supervised machine learning validation in biology. Nat Methods 18, 1122–1127 (2021). https://doi.org/10.1038/s41592-021-01205-4
Heil, B.J., Hoffman, M.M., Markowetz, F. et al. Reproducibility standards for machine learning in the life sciences. Nat Methods 18, 1132–1135 (2021). https://doi.org/10.1038/s41592-021-01256-7
Attafi OA, Clementel D, Kyritsis K, Capriotti E et al. DOME Registry: implementing community-wide recommendations for reporting supervised machine learning in biology. GigaScience. 13: giae094. (2024). https://doi.org/10.1093/gigascience/giae094